Your analysis may be perfect and your presentation very impressive, but what about your code?
For a quick throw-away analysis it may not matter.
But many projects take a lot longer. They require you to revisit and tweak your code, apply the same analysis to many different data sets and work with others. The final result needs to stand up to serious scrutiny and be reproduceable by others.
The dream
The dream is to create sleek code, which clearly expresses the steps between the problem and the solution.
There is as little code as possible, but no less.
The code is easy to understand and can be reused for other projects.
The reality
The reality is often different.
Has this ever happened to you? Your code works and you've completed your analysis. Some months later you need to do something similar. You come back to your code only to struggle to understand what it does and how it works.
Or you're working on a large project. The initial results are promising and your code works fine. Little by little it becomes harder and harder to make any changes to it. You give up and start again.
What happened? Where did it go wrong?
A trail of rubbish code mixed in with the good stuff
A complex analysis requires experimentation, different approaches, statistical models or machine learning algorithms.
You take some code off the internet and tweak it just enough to make it work.
You won't throw anything away just in case you may need it later (or am I the only?).
The result is a mixture between useful code and other stuff.
Too much to remember
The process from loading the data to the final result involves a lot of small steps, and a lot of different views and variables along the way.
You constantly scroll up and down the notebook looking for variables and their meaning.
Your brain is so busy holding on to what you've done before that there is very little space left for the anything new.
Too much code

Once you've got something working you copy the same solution to a few other cells, again and again.
You end up with a lot of duplicate code and a very long notebook.
Too fragile

You'd like to tidy your code or make some small changes, but it feels too fragile. Any change can break your code and will take hours to get working again.
Towards the dream - top tips
Here are my top tips to help you write elegant and robust data analysis code.
Clean your code .. as you go along
Before moving on to the next cell take a bit of time to clean up your code.
Remove any dead ends. And check that your notebook runs all the way through.
Clean code is easier to verify, modify and re-use.
Do it straight away. Once you have something working (e.g. load the data, or tidy up missing values), take a few minutes to tidy it up. Don't wait until the very end or it may never happen.
Keeping your code clean as you work on it will help with the next steps.
Adopt a coding standard
As you write, fix, improve or extend your code you will read it again and again.
Keep your names and layout consistent by adopting a coding standard. It cuts down on decisions and you will have less to remember.
PEP8 is the most popular Python coding standard. It was written by the creator of Python, Guido van Rossum, and two others. For more information see this article.
There are tools to help you follow PEP8, such as flake8. Your IDE (see below) may have one of those built in.
The popular Black tool will automatically tidy up your scripts.
For Jupyter Lab check out Jupyterlab Code Formatter.
For Jupyter Notebook check out Jupyter Black .
Use meaningful names
Don't use 'data' or 'df'. This is fine for examples but no use for real code.
If you can't think of a good name for something it suggests you don't understand what it is. This can trip you up later. Take a few moments to create a meaningful name.
As your data evolves, so should the name. Read the data into 'raw_population_statistics', clean them up into 'population_statistics' and group them into 'population_by_country'.
Use chaining
Some cells can only be run once. You can't delete or rename something twice. This can stop you from re-running part of your analysis.
[1]: iris = pd.read_csv('iris.csv', header=0)
[2]: iris = iris.drop('species', axis=1)
(re-run the previous cell)
[3]: iris = iris.drop('species', axis=1)
Result:
KeyError: "['species'] not found in axis"
To prevent this from happening, you could create a new data frame in each step. Your data goes from a to b to c to d to e, etc. Every time you make a change, you create a new data frame.
[1]: iris_raw = pd.read_csv('iris.csv', header=0)
[2]: iris_numbers = iris_raw.drop('species', axis=1)
(re-run the previous cell)
[3] iris_numbers = iris_raw.drop('species', axis=1)
Result: (nothing - no error)
You can now run each cell, but your meaningful names are becoming longer and longer.
Instead, use chaining. The output from the first step is used directly in the second step, etc.
iris_raw = pd.read_csv('iris.csv', header=0)
iris = (
iris_raw
.drop('species', axis=1)
.rename({
'sepal_length': 'Sepal length',
'sepal_width': 'Sepal width',
'petal_length': 'Petal lengths',
'petal_width': 'Petal width'
}, axis=1)
)
iris.head()
Result:
| Sepal length | Sepal width | Petal lengths | Petal width | |
|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 |
(etc)
Note how I used the round brackets so I can break up the line.
You don't have to write this in one go. Write the first part, run the cell to check it. Add in a bit more, and repeat until you're done. Matt Harrison has an excellent video on this.
Re-use your own code
Stop copying your code. Copying is a short term solution. In the long term this creates a big mess.
Multiple copies live in multiple places and all need the same changes when you fix or improve something. This is tedious and error prone. It is easy to forget something and end up with errors or inconsistencies.
Use Python's functions and loops instead.
Small repeat operations, for instance on multiple columns, can be done using a loop. For instance to sum quarterly results into annual results for multiple years:
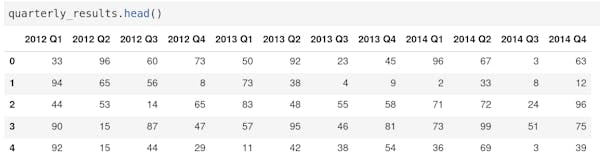
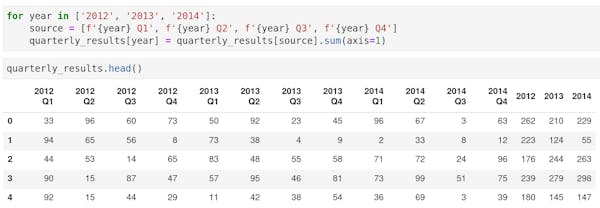
For longer operations create a Python function. Functions can be used in different places and even imported from an external script. This code does the same as the previous code but can be used on different data frames:

Result: same as above.
Use other people's code
Stop reinventing the wheel and step onto the shoulders of giants.
The Python Package Index contains over 300,000 packages. This includes 7000+ for 'pandas' and 10,000+ for 'numpy'. There are packages for machine learning, natural language processing, statistical models, computer vision and more.
Start you next machine learning project with an open source ML model.
Search GitHub for a Python library or sample code. For instance 72,000 repositories for 'pandas'.
Search on Stack Overflow or on Kaggle.
Please check the licenses to make sure you have permission to use the code.
Test your code
Detecting errors early is very helpful. It makes it much easier to fix them.
Ideally you should test all your code every time you make a change. That way you'll catch any (new) problems straight away.
Implementing a complex algorithm or model can be tricky. It will involve a lot of small steps. It is best to test each step individually.
Testing code by hand becomes time consuming, especially when you have a lot of code. It involves a lot of jumping around remembering what and how.
Save your time and brain power for the real work. Create automated tests and run them often as you need.
For testing a Python script pytest is the most popular choice. Pytest makes it very easy to write tests.
Notebooks are more complex and testing them is more tricky. nbmake helps you use pytest with your notebooks.
Know the difference between a script and a notebook
A script is a simple text file with lines of code. Scripts are great for complex code.
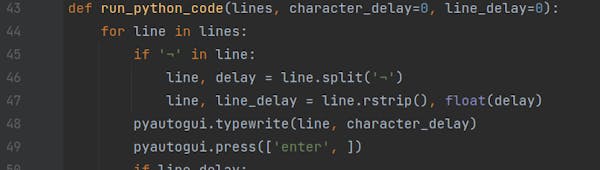
You use a text editor or Integrated Development Environment (see below) to create a script.
A notebook is a mixture of code, text and output.
Notebooks are great for experimenting and for telling the story of how you took the raw data through a process and created interesting results.
You use Jupyter or IPython to create a notebook.
Use version control
Some changes are a dead end. How do you roll those back?
Sometimes you break something by accident. But what did you change?
A version control system stores all the different versions of your code, and lets you roll back your changes.
You can see the differences between versions and work with different 'branches' for safe experimentation.
'git' is the most popular version control system for individuals and a simple way to safely share code with a team.
Python scripts are easy to track using git.
Notebooks are more complex. There isn't one single 'best' version control solution for notebooks. For many useful suggestions and real life experiences see this reddit discussion.
Keep learning
Don't copy code off the internet then just move on. You end up with a lot of code you don't understand.
Take some time to figure out how the copied code works.
Create a separate script or temporary notebook. Use the code independently from your main project. Experiment a little until you understand it. Then get back to your main code.
Make it your own. Tidying up your own code is much easier.
Make learning part of your daily routines. Listen to a Python podcast during your commute or whilst doing chores. Subscribe to an email newsletter. Challenge yourself on CheckIO, HackerRank or Kaggle.
You may discover a new library or approach which seriously improves your code.
Use the best tools .. and know how to use them
With the best tools you get better results in less time and have more fun doing it.
Use the best libraries, and learn how to use them.
If you use Jupyter learn its keyboard shortcuts.
If you write a lot of scripts, use an Integrated Development Environment (IDE).
An IDE helps you write better, more consistent, code with less effort. They make it easier to find and fix your errors.
My favourite IDE is PyCharm. I've been using it for 5+ years and wouldn't want to write any serious code without it. Another popular IDE is Microsoft's Visual Studio Code.
Finally
As a data scientist you spend most of your time with your favourite Python libraries - Pandas, Numpy, sklearn, etc. Python glues it all together for you.
Take a bit of time to become a better programmer and to write better code. Learn to write sleek and robust code and sail off into a bright future.




